Tomographic series alignment without fiducials
Tomographic reconstructions without fiducial markers are always of lesser quality than with markers. However, it is valuable in many situations where the addition of markers to a specimen may be problematic. The tilt axis and tilt angles must be known to a good accuracy. Because the tilt axis is fixed for a specific microscope installation, it can be determined by doing a fiducial-based tilt series of any arbitrary specimen (such as gold particles on carbon).
The algorithm here attempts to do subset reconstructions to improve the signal-to-noise ratio for cross-correlation. It does this iteratively to improve the alignment.
As stated in the page on preparation, the .mdoc files produced from SerialEM can be used directly in Bsoft. Alternatively, all the parameters can be set up as on the preparation page. The fastest way of aligning the tilt series is one command line:
btomaln -verb 1 -align 3,1,3 -resol 20 -edge 20,3 -lambda 2165 -out 20190117_RS1_G6_10_aln.star 20190117_RS1_G6_10.mrc.mdoc
The -align option takes three parameters:
- Number of iterations: These are iterations over the full tilt series attempting to refine the micrograph orientations
- Stopping condition: Sets the threshold to stop when the average micrograph shift difference compared to the pervious iteration drops below it.
- Number of adjacent micrographs to include in each subset reconstruction.
The -resolution option sets the high resolution limit for cross correlations
The edge option smooths the edges of the micrographs which includes erasing extraneous areas at high tilts. The parameters are the width of the edge and the standard deviation of the transition, both in pixels
The -lambda option specifies the proportionality parameter for estimating the thickness (see the section on thickness).
As with other operations, the alignment can be done from bshow. Select the "Workflow/Align micrographs" item to obtain the fllowing dialog window:
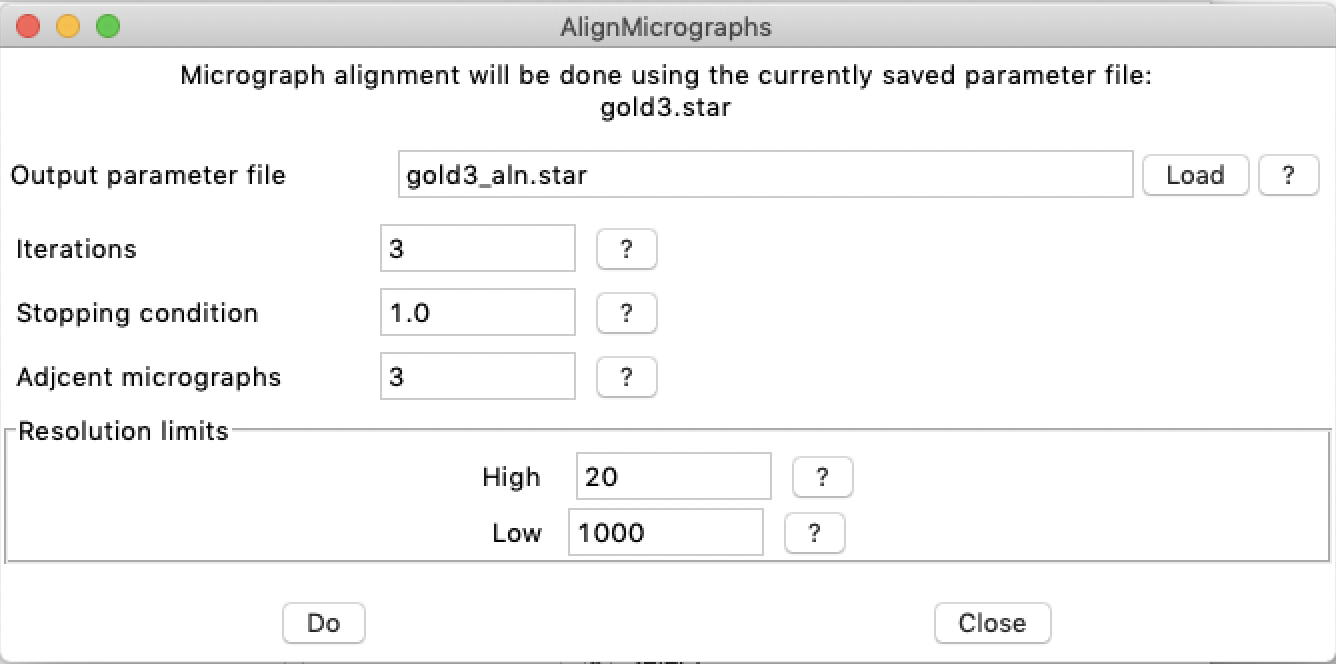
The output is written to a text window that can be saved using the "File" menu.
At this stage the results are not yet in bshow memory. Close the text window and click on the "Load" button in the "AlignMicrographs" window to load the new parameter file in memory.
Troubleshooting
Sometimes the alignment fails. While the program attempts to deal with every eventuality, it must make some assumptions regading the state of the input information. One possibility is that the direction of initial alignment biases it. To change the order in which alignment takes place, set the 180° to the tilt axis angle. Just be aware that this will change the eventual handedness of the reconstruction.
Another approach is to place markers on recognizable structures. Place one or more markers on the zero-tilt micrograph and then use the "Workflow/Generate markers from seed" menu item to produce the corresponding markers on the other micrographs. These can then be re-positioned and the alignment parameters refined (Fiducial markers).